Etiology and Epigenetics of Cancer
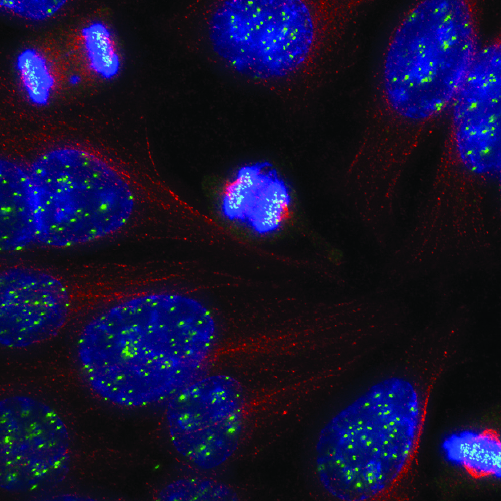
Cells must accurately regulate gene expression and maintain the integrity of their genome. Malfunctions in these processes can result in genetic diseases and cancer formation. Researchers in the division seek to understand the multiple complex molecular systems that govern chromatin structure as well as DNA transcription, RNA processing and translation, and other controls of gene expression.

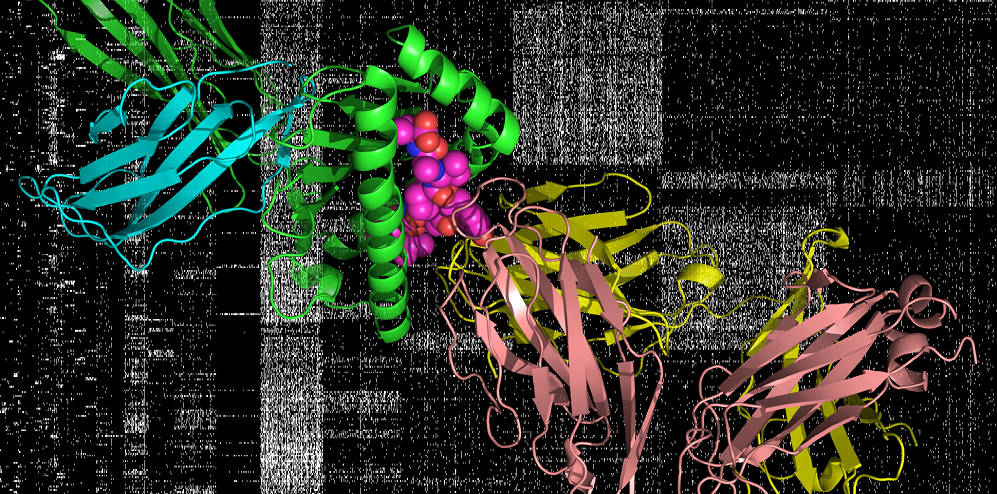
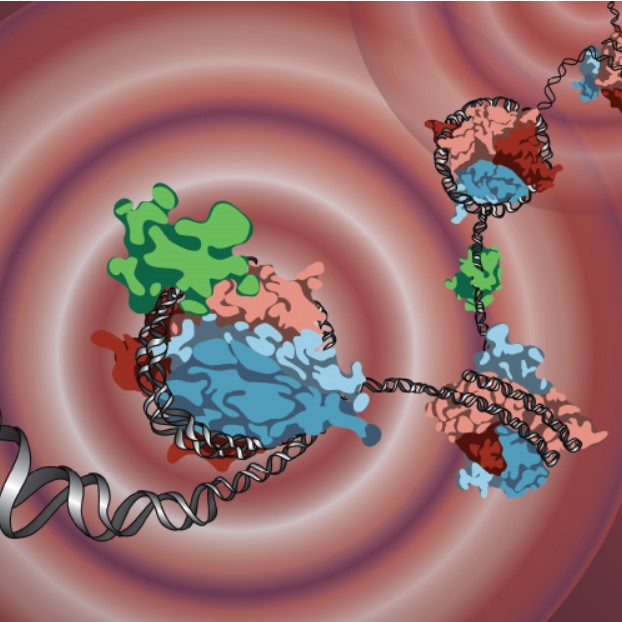
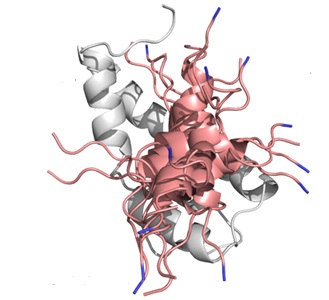
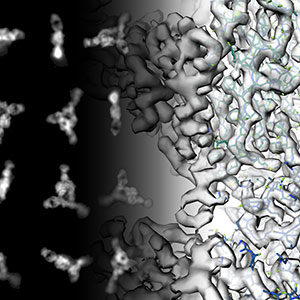
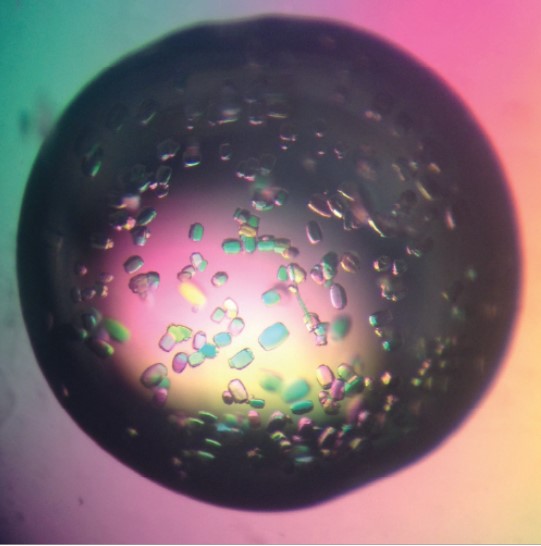
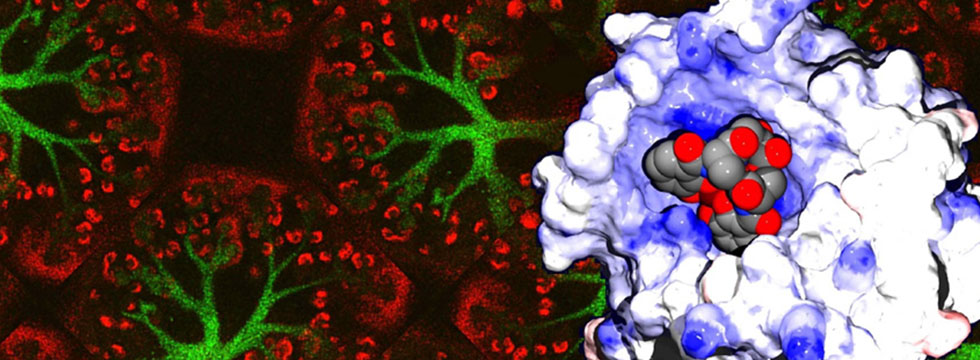
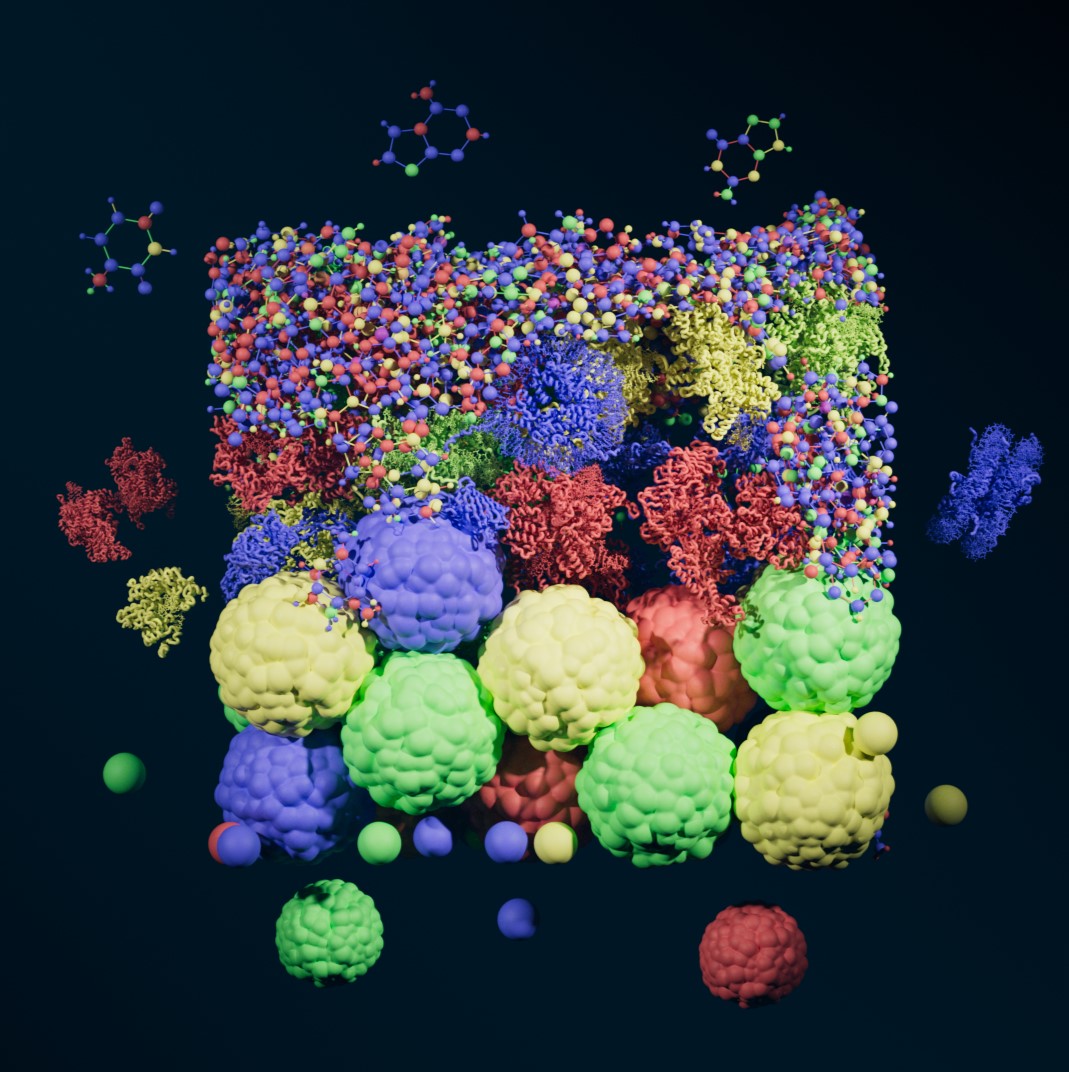
Structural and Computational Biology
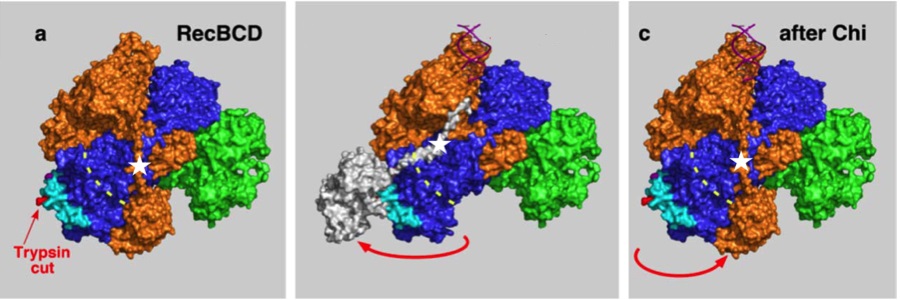
The molecular structure of proteins determines their functionality within cells and organisms. Researchers in the division are working to discover the structure of existing molecules and to synthesize new ones that might prevent or treat disease.

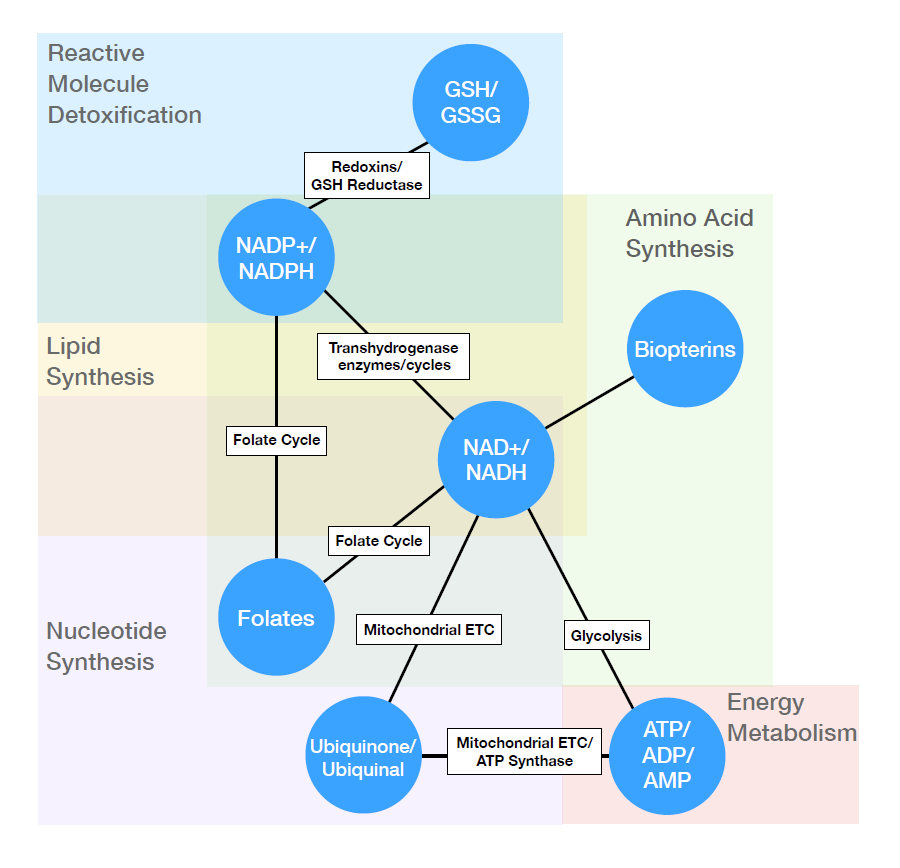
Metabolism and Stress Response
Our cells and bodies must regulate their energy and nutritional needs. This includes balancing food intake, the conversion of nutrients to the energy and building blocks for cellular processes, and the removal of unwanted proteins. Dysregulation of these process can lead to obesity, cell death and neurodegeneration, and cancer. Furthermore, the dysregulated energy needs of cancer cells can be targeted for future therapies.
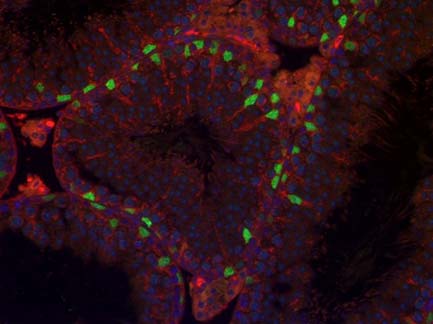
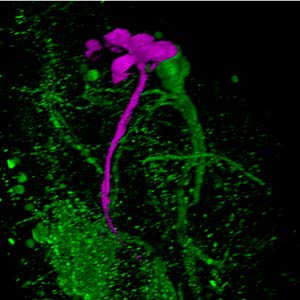
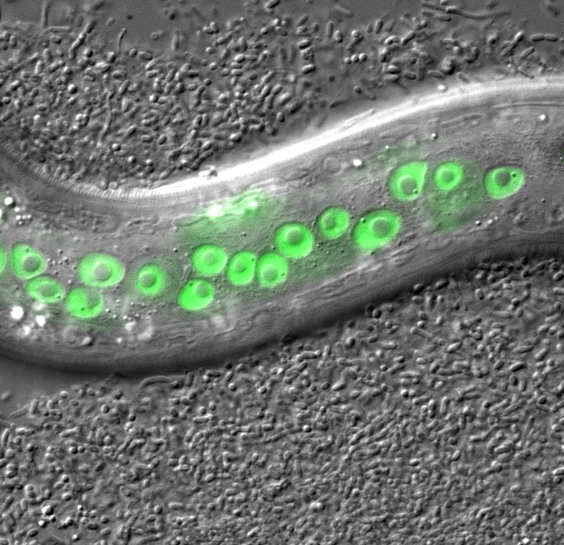

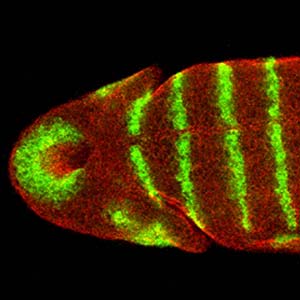
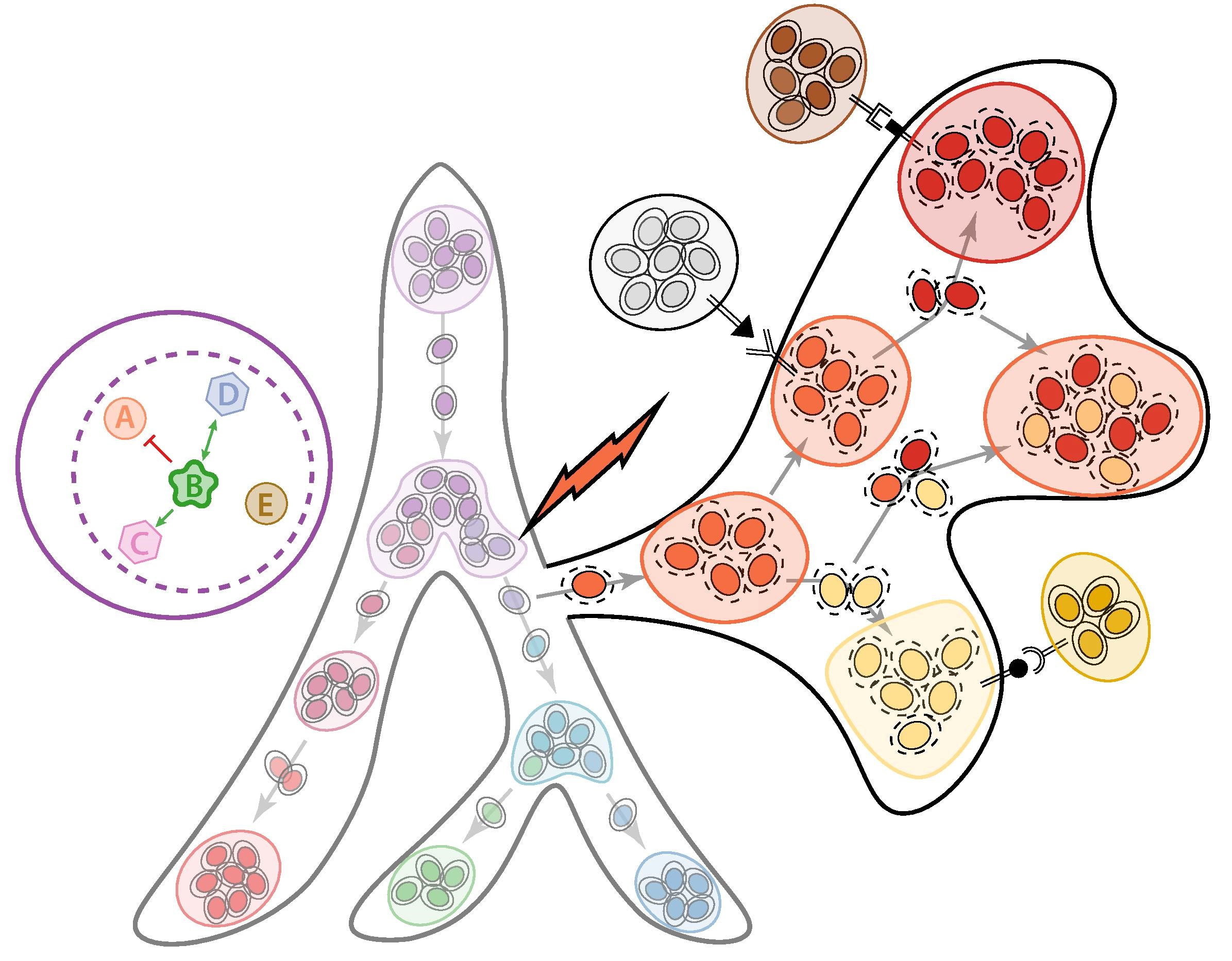
Cell and Developmental Biology
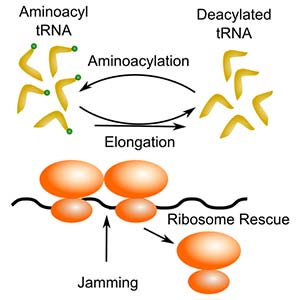
Normal cellular processes as well as environmental stressors can disrupt the genetic and structural integrity of a cell. Researchers in the division are investigating the processes by which cells recover from these stressors. Additionally, they seek to understand cellular division, metabolism and fate by answering fundamental questions about normal and abnormal cell function, development and how dysregulation of these processes contribute to disease, including cancer.





Virology, Immunology, and the Microbiota
Studying the evolution of viruses, bacteria and the immune system enables researchers to understand how these infectious agents evade immune response and develop resistance to existing therapies. Investigators also examine the symbiotic relationships among cell communities and how these affect human health. These investigations provide a deeper understanding of human immunity while revealing novel means for protecting the body from viruses like HIV and influenza.
Neuroscience
Disruptions in the proliferation, migration and signaling of cells in the brain can lead to disease and mental illness. Researchers in the division study the mechanisms behind these complex systems to better understand nervous system function and the underlying causes of neurological disorders, including brain cancers.